RNA Biology and Plasticity—Interview with Dr. John Mattick
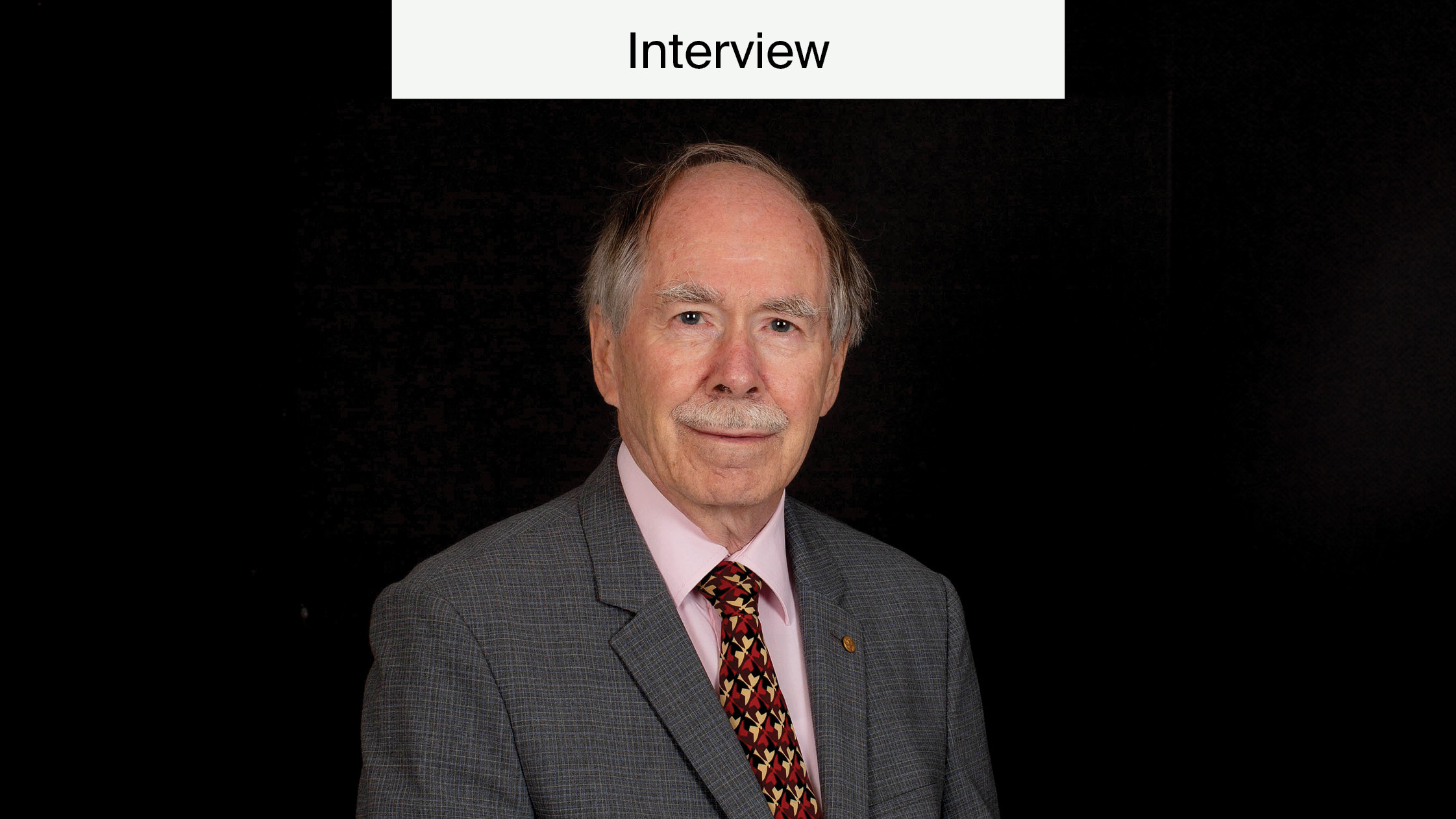
Dr. John Mattick is the Executive Director and Head of the RNA Biology and Plasticity Lab at the Garvan Institute of Medical Research. Over the past 20 years, he has made important contributions to molecular biology, such as elucidating the role of regulatory RNAs in different tissues and regions of the brain through use of advanced sequencing and bioinformatic techniques.
The human genome contains about 20,000 protein-coding genes. However, the extent of non-protein-coding DNA increases with increasing developmental and cognitive complexity. These sequences are transcribed to produce small and large non-protein-coding RNAs that guide chromatin-modifying complexes to their sites of action and thereby modulate chromatin structure and gene expression, to specify the architectural trajectories of development.
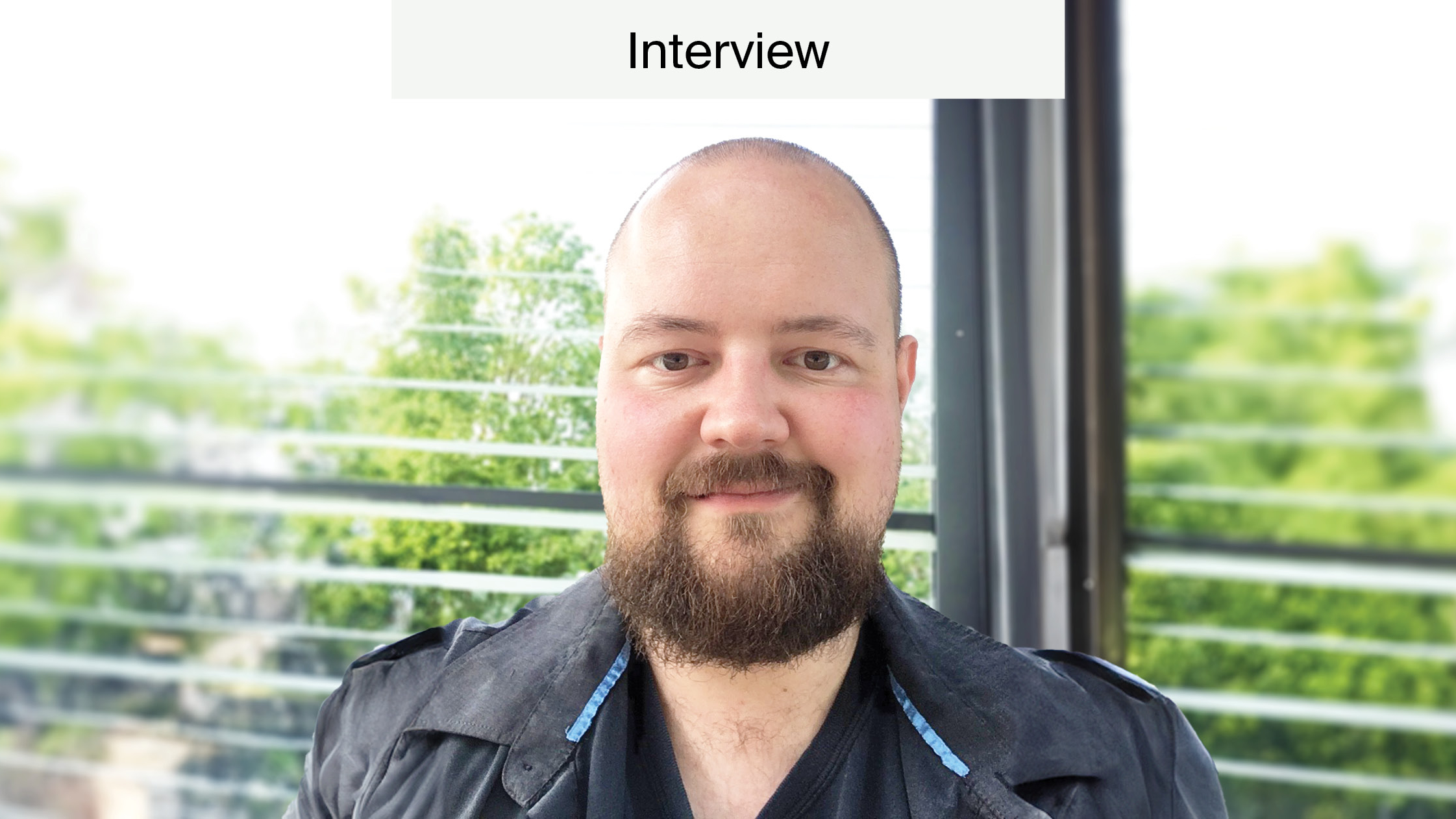
At the “Non-Coding RNAs and Epigenetics in Cancer” conference, organized by MDPI, we had the honour to interview Prof. John Mattick and find out about the latest developments and challenges in this field.
Dr. Mattick, over the past 20 years, you have pioneered a new view of the genetic programming of humans and other complex organisms. Your lab is currently focusing on characterising the expression and function of noncoding RNAs in cancer and in neurological diseases. Could you describe a few of the challenges and developments in this field?
This is a young field, we’re faced with enormous challenges in understanding the function and structure of literally tens of thousands, if not more, of regulatory RNAs that appear to be overseeing the trajectories of human development and brain function. So, the challenges within that new world are to work out how the structures of these RNAs are affecting their function and also how mutations or variations in those sequences give us pre-dispositions to complex diseases like neuropsychiatric disorders, neurodegenerative disorders and cancer. I think we are backed up on a huge endeavour; there are many of these RNAs to be analysed in terms of their biological functions. We want to understand the principles and why they work in different domains.
The thing that excites me the most is the understanding of the superimposition of plasticity on this system, through RNA editing and modification. I think that the evidence points very clearly to saying that this is the fundamental platform for the evolution of cognition and brain function. You know, the 20th century was often considered as the second-half, as being the genesis of modern molecular biology. That was just the warm-up; this is the century of biological medicine and when we understand the non-coding RNA (ncRNA), we will understand human biology.
As an experienced researcher in this field, what do you think is the most important aspect to consider when dealing with RNA research, bioinformatics and genome editing?
The most important aspect to consider in ncRNA research is the interplay between experimental data and informatic analysis. I think, in science in general, it is essential that practitioners and particularly new students coming into the field know how to handle big data sets and are confident to commission big data analysis through big genome sequencing and are also confident to do high-throughput genome editing screens to look at the functions of these RNAs. So, I think a new tool set is required, I think a new mindset is required, and I think new energy is required.
Dr John Mattick and RNA Biology
We thank Dr. John Mattick for taking the time to answer our questions. We have plenty more content on RNA biology on the blog, click here to see more.