Bayesian Statistics Meets Geometry
A common problem faced in research is when a set of data has been collected on the one hand, and a theoretical model (a set of equations with input parameters) exists on the other. We want to know firstly, whether the model describes the data well and secondly, what variables of the model give the best possible fit.
Medical data
A good example, discussed by Konukoglu, is the personalization of medical data. Specifically the data are about heart rates. And a number of models have been proposed as an explanation. In application to a single patient, however, the data is often sparse and the models are complex. It is important to know how much confidence to place in the model predictions.
Many methods exist to solve this kind of problem. An increasing number as the computational power required to crunch the data becomes cheaper. In most cases, an algorithm seeks a single value for each parameter. But, by harnessing the power of Bayesian inference, it is possible to seek a distribution of parameter values that best describe the data.
Markov chain Monte Carlo (MCMC) is an example of such a method.
It works by iteratively sampling from better and better estimates of the solution distribution to give a series of possible solution values (a Markov chain). The final answer is the distribution of these values after a certain number of iterations (the ‘burn-in’ period).
MCMC exploits the fact that the final Markov Chain must be reversible. It should converge to a point where you cannot tell if the order of points in chain sequence are swapped around at random.
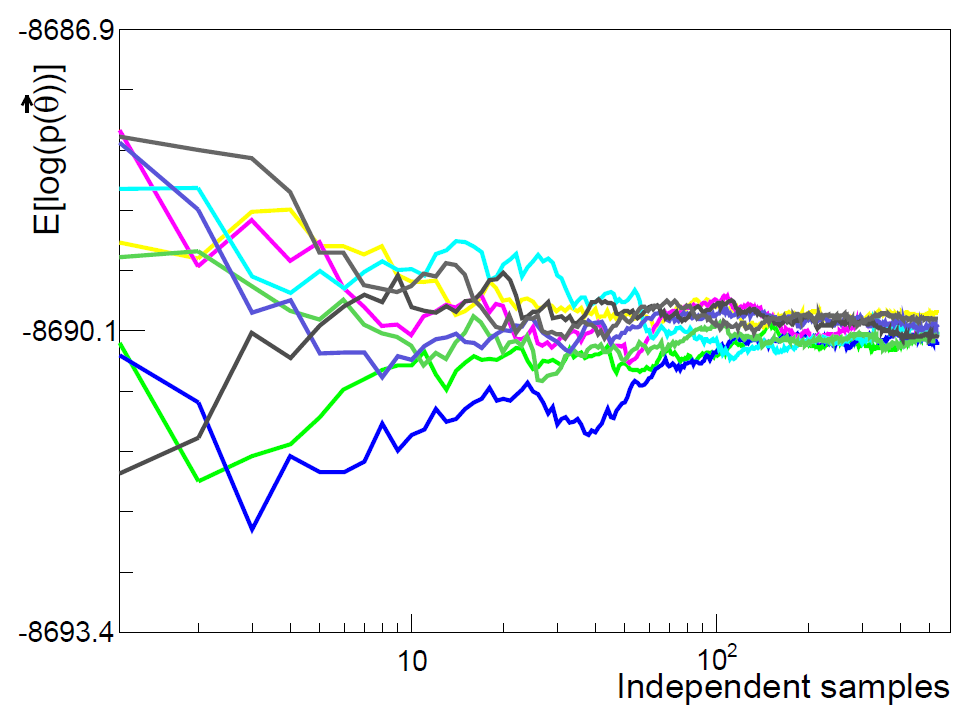
Markov Chain Monte Carlo
Some advantages of the MCMC approach are that you can easily know how precise your values are from the width of the distribution, and thus how much confidence to place in the solution. Since a distribution is broader than a single point, proposed solutions are taken from a wider area than so-called deterministic methods (e.g., those based on steepest gradients), which helps to avoid the search being trapped in a small region of the parameter space that provides only the locally best value.
To solve the problem of medical data personalization, Konukoglu et al. turned to a recent optimization of MCMC discussed by Samuel Livingstone and Mark Girolami in a paper in Entropy.
They present the Metropolis-Hastings MCMC algorithm and offer some ways to improve it for particularly difficult cases. They build on their earlier work in which they presented a method that makes use of the geometry of the parameter space to improve the chance of Markov chain conversion.
Drawbacks of Markov chain Monte Carlo
A major drawback of MCMC is that, unlike a deterministic algorithm, there is no sure-fire way to know when you should stop or how long convergence will take. The Markov chain may even never converge. Practitioners try to modify the properties of the input distribution (known as the proposal kernel) to give a series of outputs that will quickly converge. The optimum proposal kernel is dependent on the shape of the final distribution.
To deal with these issues, Livingstone and Girolami show that the use of Langevin diffusion in the decision step can lead to an increase of the optimum acceptance rate from 0.234 to 0.574. This saves valuable computational time. The resulting algorithm, is known as the Metropolis-adjusted Langevin algorithm (MALA).
Taking this further, they incorporate ideas from Riemannian geometry. This is a branch of mathematics dealing with smooth manifolds including how to define a metric: a concept of distance. An intelligent choice of metric can make points that were previously far apart become closer.
Applied to MCMC, this means that points from which the Markov chain might diverge can be moved closer to ones from which it will converge, increasing the chance of a positive and quick result. The new algorithm is termed ‘MALA on a manifold’.
Mapping MALA
A mapping from one metric to another can be given by a function G and clearly the choice of G is critical. Livingstone and Girolami show how the Fisher information and Hessian can be used.
Some interesting issues remain outstanding, such as how to avoid unwanted properties of G. For example, the best choice should be ‘positive definite’, essentially meaning that distances between two points shouldn’t become zero or negative, which is not always the case for the Hessian. Also, the further complexity involved means that the manifold method is also not necessary for many cases. How does one go about deciding when to use it?
The interest of the MALA on a manifold method lies not just in widening the applicability of MCMC, but also in the steps forward that can be made by combining two very different fields of mathematical science: Bayesian statistics and geometry. It is a fine example of the benefits of a multidisciplinary approach.
Authors writing about Markov chains

Samuel Livingstone is a PhD student in Statistical Science at University College London, under the supervision of Professor Mark Girolami and Dr. Alexandros Beskos. He is interested in the properties of estimators constructed using Markov chain Monte Carlo methods, and in understanding how ideas from Riemannian Geometry can alter these properties. Before his PhD he studied for an MSc in Statistics at Lancaster University, after a short previous career in consultancy.

Mark Girolami is a Professor of Statistics at the University of Warwick. He holds an honorary professorship in Computer Science at Warwick. Hi is an EPSRC Established Career Fellow (2012–2017) and previously an EPSRC Advanced Research Fellow (2007–2012). He is also honorary Professor of Statistics at University College London, is the Director of the EPSRC funded Research Network on Computational Statistics and Machine Learning and in 2011 was elected to the Fellowship of the Royal Society of Edinburgh when he was also awarded a Royal Society Wolfson Research Merit Award.
References for further reading about Markov chains
Konukoglu, E.; Relan, J.; Cilingir, U.; Menze, B.H.; Chinchapatnam, P.; Jadidi, A.; Cochet, H.; Hocini, M.; Delingette, H.; Jaïs, P.; Haïssaguerre, M.; Ayache, N.; Sermesant, M. Efficient probabilistic model personalization integrating uncertainty on data and parameters: Application to Eikonal-Diffusion models in cardiac electrophysiology. Prog. Biophys. Mol. Bio. 2011, 107, 134–146; doi:10.1016/j.pbiomolbio.2011.07.002.
Billard, J.; Mayet, F.; Santos, D. Markov Chain Monte Carlo analysis to constrain Dark Matter properties with directional detection. ArXiv preprint, arXiv:1012.3960 [astro-ph.CO]; doi:10.1103/PhysRevD.83.075002.
Livingstone, S.; Girolami, M. Information-Geometric Markov chain Monte Carlo methods using diffusions. Entropy 2014, 16, 3074–3102; doi:10.3390/e16063074.
Metropolis, N.; Rosenbluth, A.W.; Rosenbluth, M.N.; Teller, A.H. Equation of state calculations by fast computing machines. J. Chem. Phys. 1953, 21, 1987–1092.
Hastings, W.K. Monte-Carlo sampling methods using Markov chains and their applications. Biometrika 1970, 57, 97–109; doi:10.1093/biomet/57.1.97.
Girolami, M.; Ben Calderhead, B. Riemann manifold Langevin and Hamiltonian Monte Carlo methods. J. R. Statist. Soc. B 2011, 73, 1–37; doi:10.1111/j.1467-9868.2010.00765.x